HSP Examples: DNA
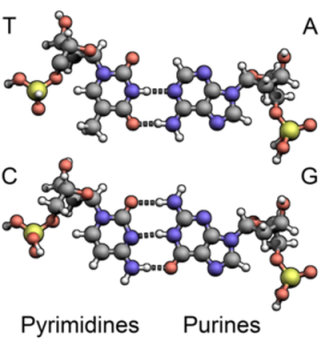 It might seem odd that DNA can have an HSP value. Surely all those phosphates take DNA way outside any sensible HSP zone?
It might seem odd that DNA can have an HSP value. Surely all those phosphates take DNA way outside any sensible HSP zone?
That's true, but the core of DNA is the base pairs and via many different paths it becomes clear that the HSP of DNA as a whole is in the range of [19, 20, 11] and it is no coincidence that the HSP average of the four DNA bases is [19.8, 12.3, 12.2] - in other words, the HSP of DNA is dominated by the bases but moved in a more hydrophilic direction via the ribose phosphate portion.
What can be done with such values? A lot!
First, there is a reasonable hypothesis that for a molecule to be cytotoxic it should, at a minimum, be "soluble" within the DNA core. Here is a list of 7 well-known cytotoxic drugs which, for obvious reasons, are often chemotherapy agents. Their HSP Distances from a nominal [19.8, 12.3, 12.2] DNA are shown:
| Chemical | δD | δP | δH | Distance |
|---|---|---|---|---|
| Fluorouracil | 18 | 11.7 | 11.6 | 3.7 |
| Gemcitabine | 19 | 12.6 | 15.5 | 3.7 |
| Cyclophosphamide | 17.5 | 11.9 | 12.6 | 4.6 |
| Ifosfamide | 17.5 | 11.9 | 9.8 | 5.2 |
| Methotrexate | 18 | 10.2 | 14.2 | 4.6 |
| Etoposide | 20 | 7.5 | 12.5 | 4.8 |
| Paclitaxel (Taxol) | 18 | 6.6 | 9.8 | 7.2 |
All these molecules are within striking distance of the DNA bases. When such a molecule resides in the vicinity of the bases a physical block is formed to the normal DNA functions such as replication. In passing it might be noted that the HSP Distances of the DNA bases from water is large (>8) and it is well-known that they are not water soluble. The implications of this important point are discussed below.
These data, along with others described in the HSPiP eBook suggest that biologists are missing a trick when they ignore the "solubility" aspects of DNA.
But there is more. If you want to rapidly screen, say, a tumour for a specific DNA sequence in order to identify its particular type, one way is to use FISH hybridisation. For this the DNA double helix needs to be gently separated into individual strands which then have a chance to hybridise (if a close match) with fluorescent-labelled probe DNA sequences.
For many years the standard way to do this was by adding formamide to the aqueous solution containing the DNA. Unfortunately the hybridisation with formamide is very slow. Could a better FISH solvent be found? Charles, working with the DAKO company in Denmark naturally turned to HSP to help in the search. A list of potential candidates was found starting with the HSP sphere with “good” solvents being formamide and DMSO. Several of these candidates were tested with reasonable success and a new, improved HSP sphere could be generated with these data. Among these, ethylene carbonate (EC) was found most advantageous because of its water solubility. As expected, a final calculation provided the answer why EC works so well. Taking the [19, 20, 11] value for DNA in water and using the classical approach that two bad solvents can make a good solvent, then a blend of 50:50 Ethylene Carbonate [19.4, 21.7, 5.1] and "structured" water [18.1, 17.1, 16.9] gives an HSP of [18.7, 19.3, 11.0], a near-perfect match for the DNA. Experiments confirmed this. Rather than taking 1 day for a FISH hybridisation, it can be done in 1 hour. In the case of an urgent diagnosis for a critically ill patient this can be (and has been) a matter of life and death.
What else might be accomplished via these insights into DNA HSP? That's up to those who choose to take these ideas and test them in other settings. Charles is confident that it is only a matter of time before new applications for HSP in DNA-related work will appear.
Finally, it is worth pointing out that it is not hydrogen bonding that “binds” the helices with each other (though of course it is the cause of the specificity of base-pair binding). The hydrogen bonding parameter for DNA is clearly the lowest of the three HSP, and as pointed out above, the HSP Distance to water is large. It is the essentially zero water solubility of the DNA bases as pure compounds that drives them together into the hydrophobic core of the DNA. This hydrophobic driving energy is large, and the energy of the hydrogen bonds that establish the specific base pairings is small in comparison.